TAPS+ Positive Methylation Library Construction Solution
In the native mammalian genome, unmethylated cytosines account for over 95%, while methylated cytosines (C) make up less than 5%. Conventional BS/EM-seq methods extensively convert unmethylated C, drastically altering the sequence information of the reference genome.
As a next-generation bisulfite-free methylation sequencing technology, TAPS relies on TET enzyme oxidation and pyridine borane reduction reactions. It only site-specifically modifies a small number of modified sites, positively converting methylated C to T while fully preserving unmethylated C.
This solution effectively overcomes limitations of conventional methods such as sequence damage, information loss, and alignment difficulties. It better maintains genome integrity, base complexity, and stable GC content, significantly improving sequencing quality, alignment efficiency, and bioinformatics analysis throughput. Meanwhile, it fully retains variant information such as SNV and Indel, enabling simultaneous analysis of methylation profiles and genomic variations, and is widely adaptable to various methylation research scenarios.
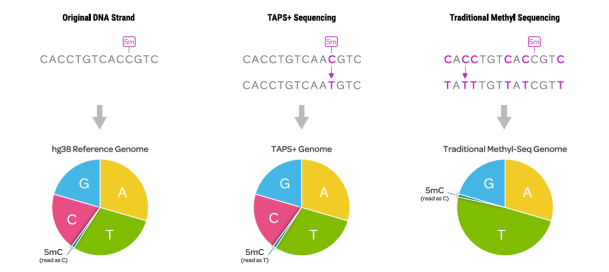
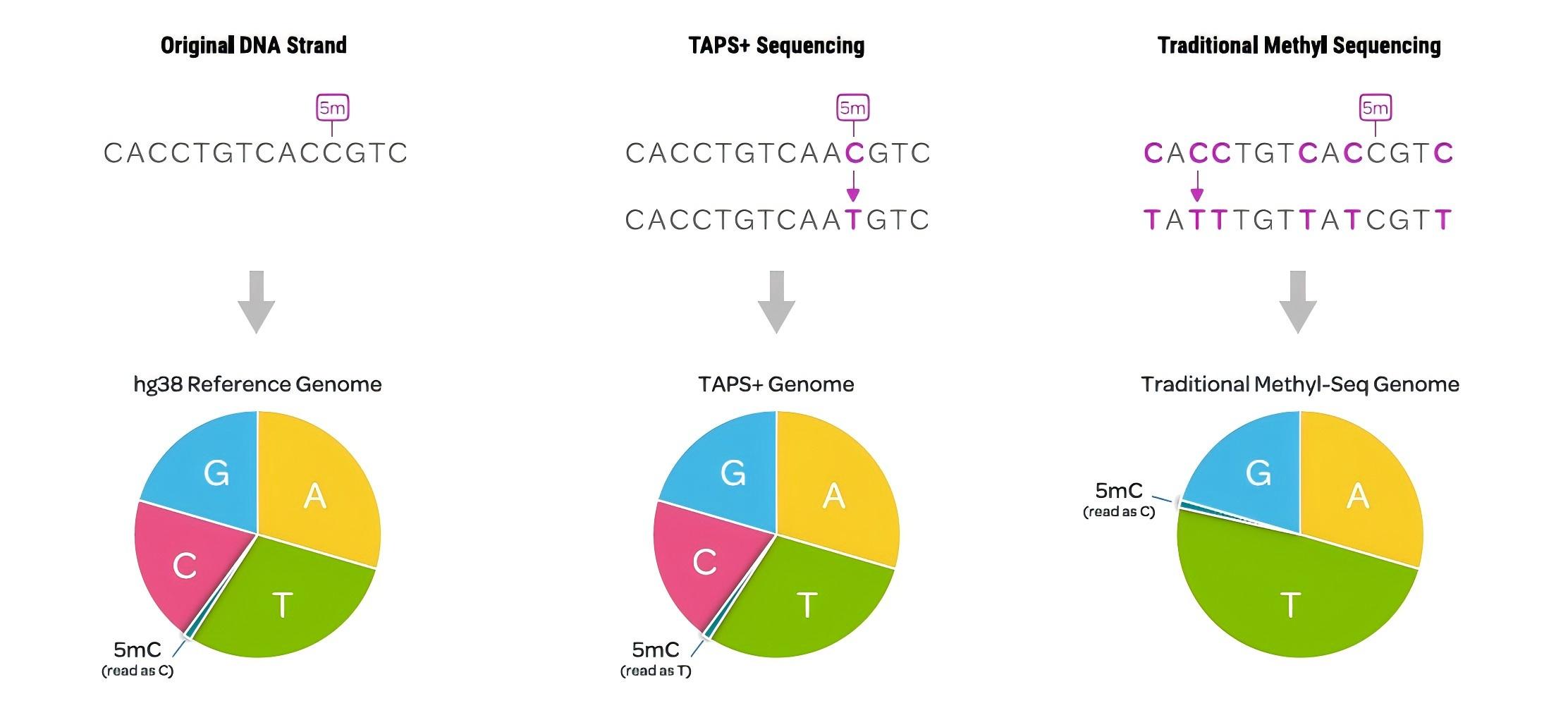
Figure 1. Compared with conventional methylation sequencing, TAPS preserves the complexity of the four natural bases and directly reads 5mC signals.

iGenetech TAPS+ Library Capture Solution
Building on profound expertise in gene capture technology, proprietary probe design algorithms, and a mature experimental system, iGenetech introduces a positive-strand capture solution compatible with Watchmaker Genomics TAPS+ libraries.
On the basis of its two established methylation capture platforms — BisCap® (capture-after-conversion) and MethCap® (conversion-after-capture) — iGenetech has added the TargetSeq One® universal hybrid capture system, which is directly compatible with the TAPS+ library solution, further expanding methylation detection workflows.
Leveraging the high enrichment efficiency of targeted capture, the solution supports flexible selection of target regions and rapid customization of high-performance probes. It enables simultaneous and accurate detection of sample methylation levels and genetic variations, providing one-stop multi-dimensional analysis of epigenetics and molecular variants.
Protocol Introduction and Performance Data
1、TAPS+ libraries can be directly used with universal hybrid capture kits and share a unified reagent system with non-methylation capture panels, greatly simplifying kit and experimental workflow management. It achieves more efficient and uniform enrichment of target regions with a simple and robust workflow.


Figure 2. Capture performance test. Prepare library with 100 ng NA12878 TAPS+ (test library provided by Watchmaker Genomics), using 1.7 Mb panel with TargetSeq One® Hyb & Wash Kit v3.0 for capture, sequenced on SURFSeq 5000 with 2G raw bases, ~ 440× depth.
2、Dual-state design plus Waston-Crick strand probe strategy ensures efficient capture of target regions while maintaining detection performance. Compared with probe design based on direct reference genome, this approach ensures effective capture of fragments with diverse methylation states and multiple CG-dense regions.

Figure 3. TAPS dual-state probe design for methylation detection.
3、97.6% conversion efficiency enables efficient enrichment of targeted fragments and precise capture of key methylation sites, delivering stable and reliable data.

Figure 4. Positive conversion efficiency evaluated using internal control λ-DNA. Prepare library with 100 ng NA12878 TAPS+ (test libraries provided by Watchmaker Genomics), using a 1.7 Mb panel with TargetSeq One® Hyb & Wash Kit v3.0 for capture, sequenced on SURFSeq 5000 with 2G raw bases, ~ 440× depth.
4、Methylation levels between technical replicates are highly consistent, with low batch variation and excellent reproducibility, enabling stable and accurate detection of sample methylation levels.

Figure 5. Stability analysis across technical replicates. Methylation consistency analysis of data from different technical replicates of TAPS+ library capture.
5、Highly consistent with WGBS “Golden Standard”methylation levels, showing excellent result concordance, accurately restoring epigenetic features and enabling reliable detection of methylation levels.

Figure 6. Methylation level consistency analysis. Methylation levels in target regions from TAPS+ library capture compared with WGBS results.
Product Information
| | |
| TargetSeq One® Hyb & Wash Kit v3.0 with Eco Universal Blocking Oligo* | | |
| TargetSeq® Cap Beads & Nuclease-Free Water | | |
TAPS+ products are from Watchmaker Genomics. For technical and purchasing inquiries, please contact their China representative:
Mr. Liang
Tel: 13693619015
Email: Haitao.liang@watchmakergenomics.cn
For iGenetech panel customization and hybrid capture reagents, technical support and custom requests, please contact our Sales Director:
Mr. Lang
Tel: 18701592757
Email: xianyong.lang@igenetech.com
About iGenetech
iGenetech is a China-based high-tech enterprise focused on developing and supplying target gene “reading” and “writing” technology solutions. It owns three core proprietary technology platforms: NGS probe hybridization, multiplex PCR, and high-throughput DNA synthesis.
iGenetech is certified under ISO 13485 and ISO 9001 quality management systems, equipped with a ~ 2,500 ㎡ GMP-compliant manufacturing facility and testing laboratory. It provides customized gene capture products, targeted sequencing laboratory solutions, NGS rapid detection services, and large-scale DNA synthesis services for healthcare, agriculture, microbiology, academic research, and other fields.
Website: www.igenetech.com
About Watchmaker Genomics
Watchmaker Genomics is a life science company dedicated to pioneering high-performance enzyme engineering to advance genomics, molecular diagnostics, and synthetic biology. Using advanced directed evolution and precision protein design, Watchmaker develops molecular enzymes with industry-leading performance, improving accuracy, sensitivity, and efficiency in next-generation sequencing (NGS) and molecular diagnostic (MDx) workflows.
Headquartered in Boulder, Colorado, Watchmaker collaborates with top global researchers, diagnostic developers, and biotechnology companies to expand the boundaries of molecular biology.
Website: watchmakergenomics.com

 CN
CN


